Going from a research idea to experiments is fundamental. But this step is typically glossed over with little explicit advice. In academia, the graduate student is often left toiling away—fragmented code, various notes and LaTeX write-ups scattered around. New projects often result in entirely new code bases, and if they do rely on past code, are difficult to properly extend to these new projects.
Motivated by this, I thought it’d be useful to outline the steps I personally take in going from research idea to experimentation, and how that then improves my research understanding so I can revise the idea. This process is crucial: given an initial idea, all my time is spent on this process; and for me at least, the experiments are key to learning about and solving problems that I couldn’t predict otherwise.1
Finding the Right Problem
Before working on a project, it’s necessary to decide how ideas might jumpstart into something more official. Sometimes it’s as simple as having a mentor suggest a project to work on; or tackling a specific data set or applied problem; or having a conversation with a frequent collaborator and then striking up a useful problem to work on together. More often, I find that research is a result of a long chain of ideas which were continually iterated upon—through frequent conversations, recent work, longer term readings of subjects I’m unfamiliar with (e.g., Pearl (2000)), and favorite papers I like to revisit (e.g., Wainwright & Jordan (2008), Neal (1994)).

One technique I’ve found immensely helpful is to maintain a single master document.2 It does a few things.
First, it has a bulleted list of all ideas, problems, and topics that I’d like to think more carefully about (Section 1.3 in the figure). Sometimes they’re as high-level as “Bayesian/generative approaches to reinforcement learning” or “addressing fairness in machine learning”; or they’re as specific as “Inference networks to handle memory complexity in EP” or “analysis of size-biased vs symmetric Dirichlet priors.”. I try to keep the list succinct: subsequent sections go in depth on a particular entry (Section 2+ in the figure).
Second, the list of ideas is sorted according to what I’d like to work on next. This guides me to understand the general direction of my research beyond present work. I can continually revise my priorities according to whether I think the direction aligns with my broader research vision, and if I think the direction is necessarily impactful for the community at large. Importantly, the list isn’t just about the next publishable idea to work on, but generally what things I’d like to learn about next. This contributes long-term in finding important problems and arriving at simple or novel solutions.
Every so often, I revisit the list, resorting things, adding things, deleting things. Eventually I might elaborate upon an idea enough that it becomes a formal paper. In general, I’ve found that this process of iterating upon ideas within one location (and one format) makes the transition to formal paper-writing and experiments to be a fluid experience.
Managing Papers
Good research requires reading a lot of papers. Without a good way of organizing your readings, you can easily get overwhelmed by the field’s hurried pace. (These past weeks have been especially notorious in trying to catch up on the slew of NIPS submissions posted to arXiv.)
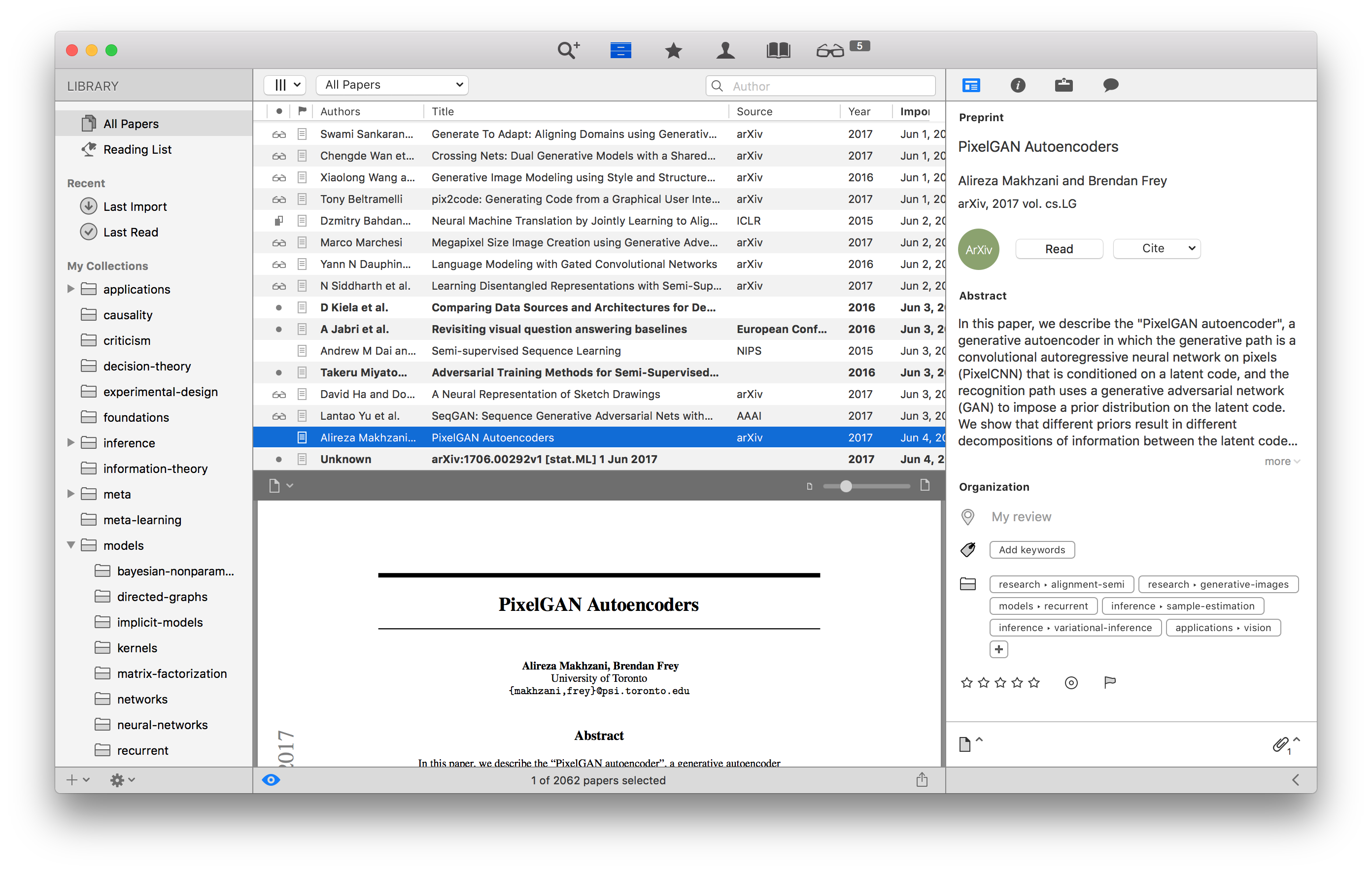
I’ve experimented with a lot of approaches to this, and ultimately I’ve arrived at the Papers app which I highly recommend.3
The most fundamental utility in a good management system is a centralized repository which can be referenced back to. The advantage of having one location for this cannot be underestimated, whether it be 8 page conference papers, journal papers, surveys, or even textbooks. Moreover, Papers is a nice tool for actually reading PDFs, and it conveniently syncs across devices as I read and star things on my tablet or laptop. As I cite papers when I write, I can go back to Papers and get the corresponding BibTeX file and citekey.
I personally enjoy taking painstaking effort in organizing papers. In
the screenshot above, I have a sprawling list of topics as paper tags.
These range from applications, models, inference (each with
subtags), and there are also miscellaneous topics such as
information-theory and experimental-design. An important
collection not seen in the screenshot is a tag called research,
which I bin all papers relevant to a particular research topic into.
For example, the PixelGAN paper
presently highlighted is tagged into two topics I’ve currently been
thinking a lot about—these are sorted into research→alignment-semi
and research→generative-images.
Managing a Project
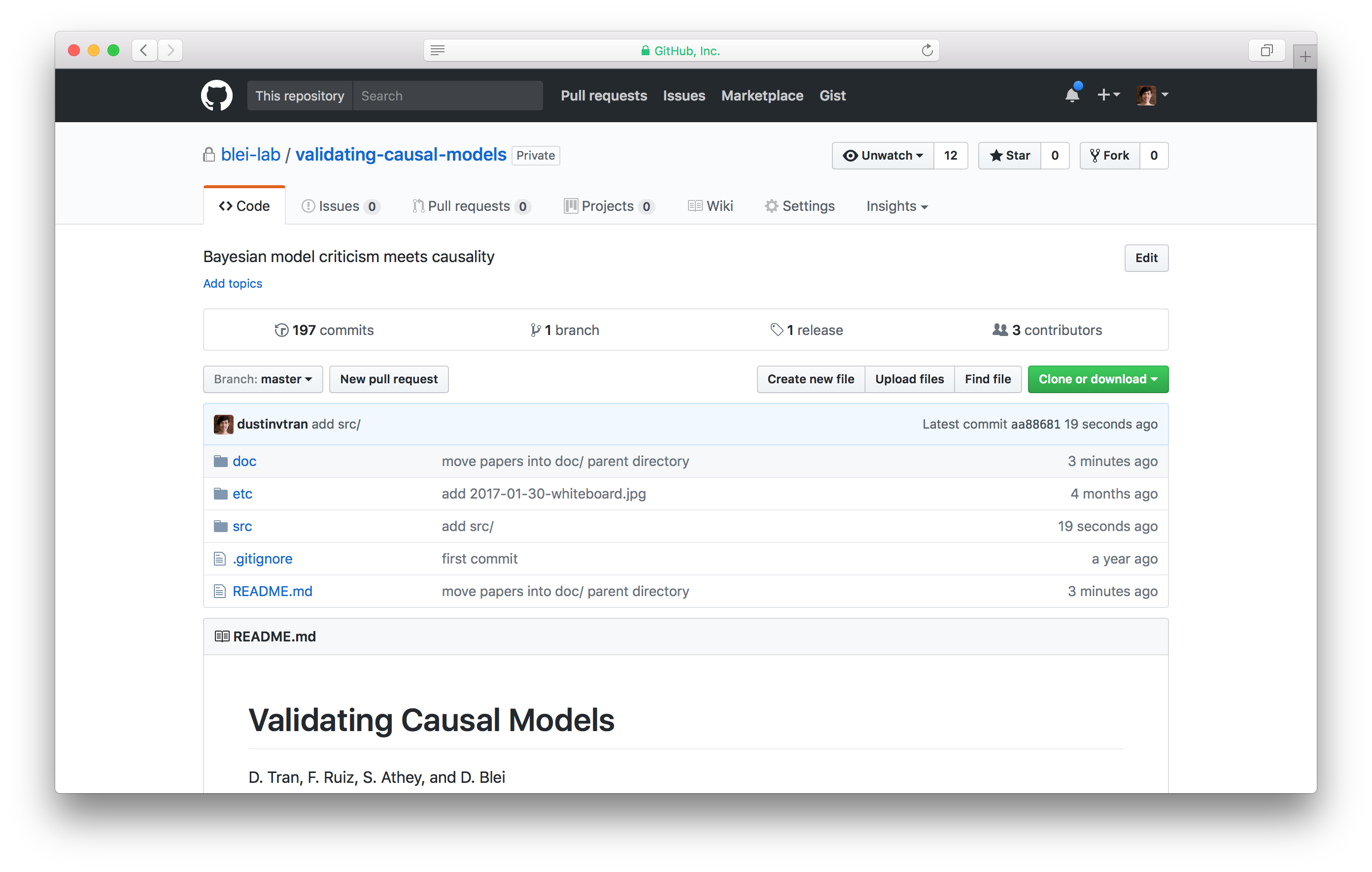
I like to maintain one research project in one Github repository. They’re useful not only for tracking code but also in tracking general research progress, paper writing, and tying others in for collaboration. How Github repositories are organized is a frequent pain point. I like the following structure, based originally from Dave Blei’s preferred one:
-- doc/
-- 2017-nips/
-- preamble/
-- img/
-- main.pdf
-- main.tex
-- introduction.tex
-- etc/
-- 2017-03-25-whiteboard.jpg
-- 2017-04-03-whiteboard.jpg
-- 2017-04-06-dustin-comments.md
-- 2017-04-08-dave-comments.pdf
-- src/
-- checkpoints/
-- codebase/
-- log/
-- out/
-- script1.py
-- script2.py
-- README.md
README.md maintains a list of todo’s, both for myself and
collaborators. This makes it transparent how to keep moving forward
and what’s blocking the work.
doc/ contains all write-ups. Each subdirectory corresponds to a
particular conference or journal submission, with main.tex being the
primary document and individual sections written in separate files
such as introduction.tex. Keeping one section per file makes
it easy for multiple people to work on separate sections
simultaneously and avoid merge conflicts. Some people prefer to write
the full paper after major experiments are complete. I personally like to
write a paper more as a summary of the current ideas and, as with the
idea itself, it is continually revised as experiments proceed.
etc/ is a dump of everything not relevant to other directories. I
typically use it to store pictures of whiteboards during conversations
about the project. Or sometimes as I’m just going about my day-to-day,
I’m struck with a bunch of ideas and so I dump them into a Markdown
document. It’s also a convenient location to handle various
commentaries about the work, such as general feedback or paper
markups from collaborators.
src/ is where all code is written. Runnable scripts are written
directly in src/, and classes and utilities are written in
codebase/. I’ll elaborate on these next. (The other three are
directories outputted from scripts, which I’ll also elaborate upon.)
Writing Code
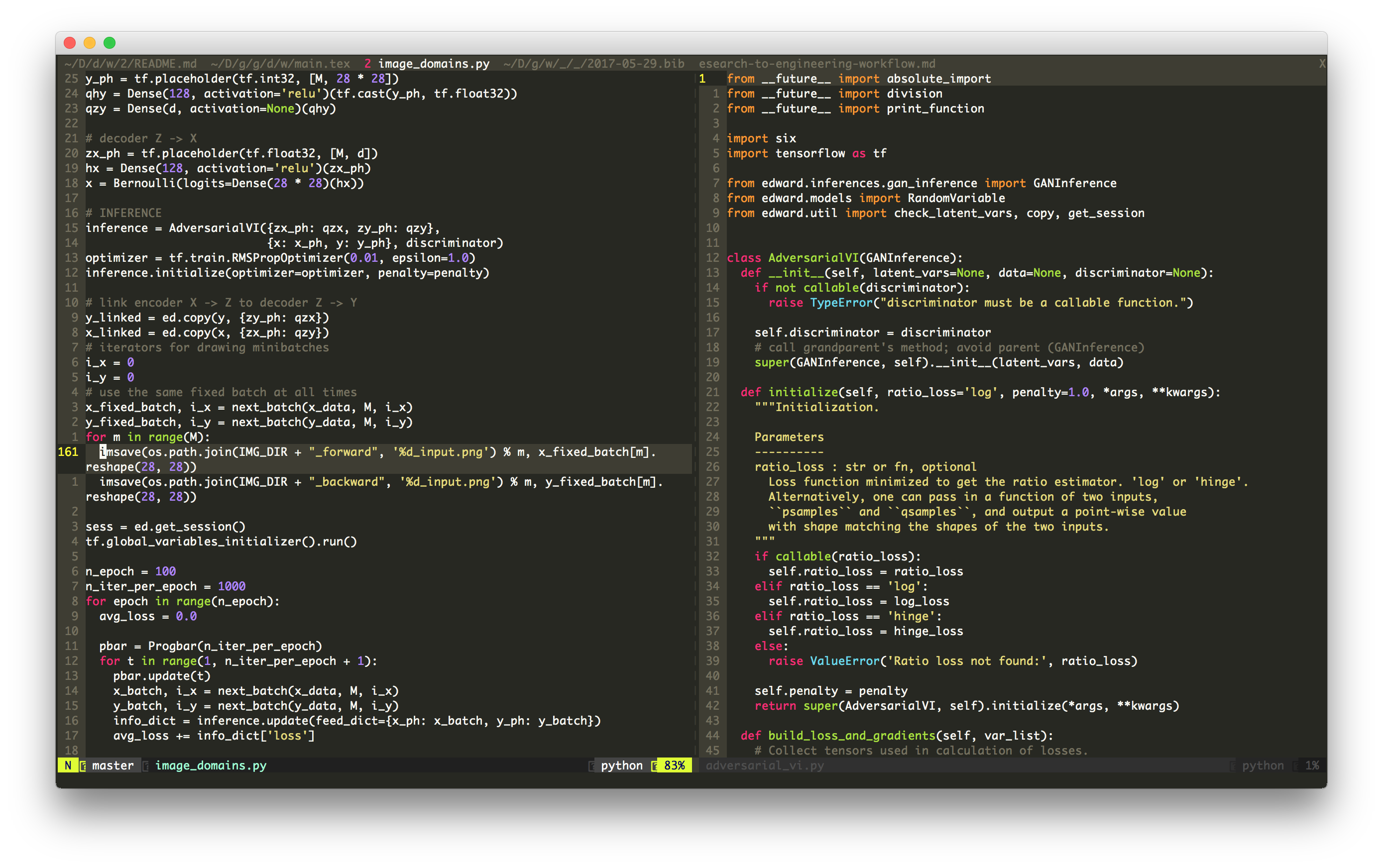
Any code I write now uses Edward. I find it to be the best framework for quickly experimenting with modern probabilistic models and algorithms.
On a conceptual level, Edward’s appealing because the language explicitly follows the math: the model’s generative process translates to specific lines of Edward code; then the proposed algorithm translates to the next lines; etc. This clean translation avoids future abstraction headaches when trying to extend the code with natural research questions: for example, what if I used a different prior, or tweaked the gradient estimator, or tried a different neural net architecture, or applied the method on larger scale data sets?
On a practical level, I most benefit from Edward by building off
pre-existing model examples
(in edward/examples/ or edward/notebooks/),
and then adapting it to my problem.
If I am also implementing a new
algorithm, I take a pre-existing algorithm’s source
code (in edward/inferences/),
paste it as a new file in my research project’s codebase/ directory,
and then I tweak it. This process makes it really easy to start
afresh—beginning from templates and avoiding low-level details.
When writing code, I always follow PEP8 (I particularly like the
pep8 package), and I try
to separate individual scripts from the class and function definitions shared
across scripts; the latter is placed inside codebase/ and then imported.
Maintaining code quality from the beginning is always a good
investment, and I find this process scales well as the code gets
increasingly more complicated and worked on with others.
On Jupyter notebooks. Many people use Jupyter notebooks as a method for interactive code development, and as an easy way to embed visualizations and LaTeX. I personally haven’t found success in integrating it into my workflow. I like to just write all my code down in a Python script and then run the script. But I can see why others like the interactivity.
Managing Experiments
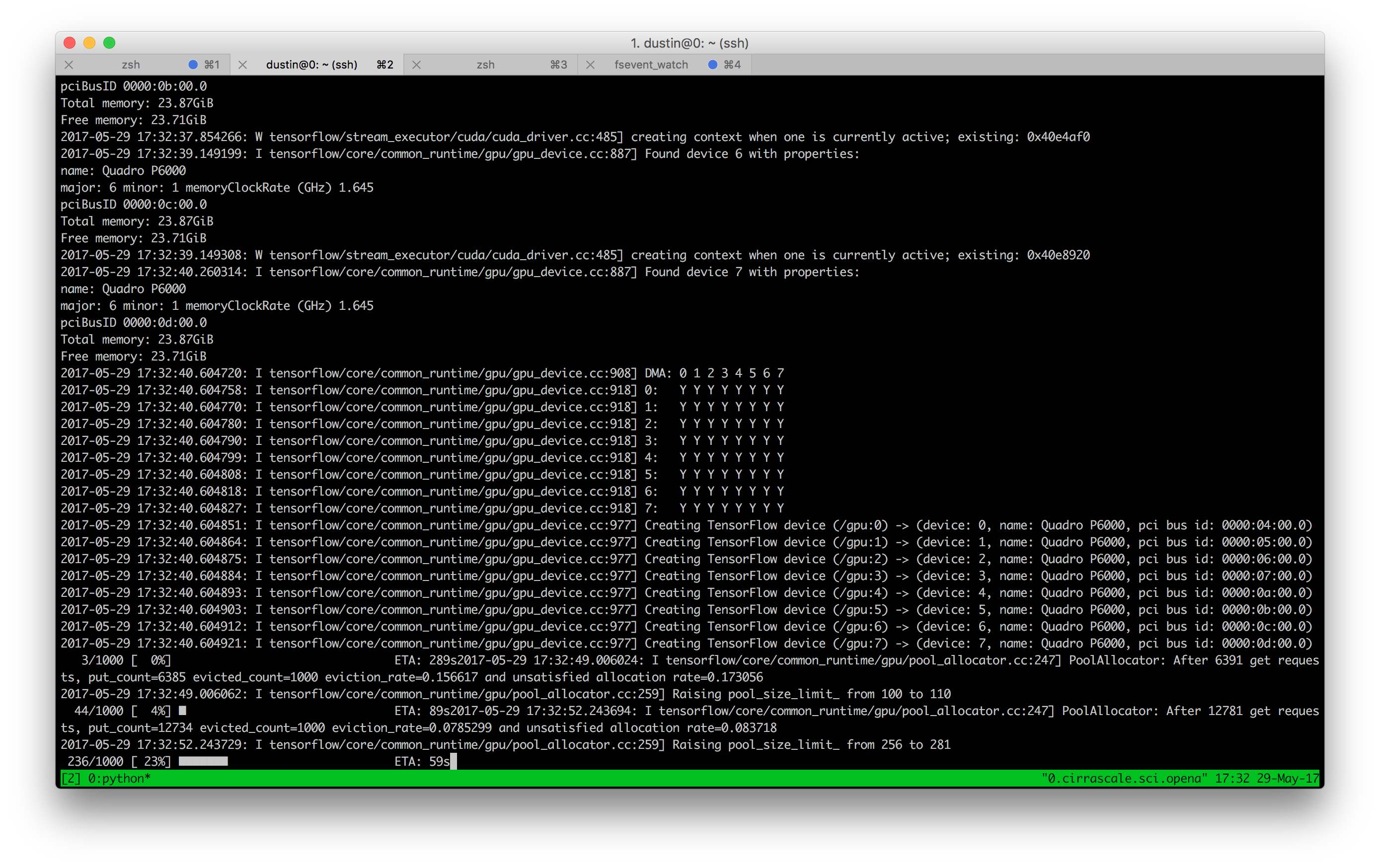
Investing in a good workstation or cloud service is a must. Features such as GPUs should basically be a given with their wide availability, and one should have access to running many jobs in parallel.
After I finish writing a script on my local computer, my typical workflow is:
- Run
rsyncto synchronize my local computer’s Github repository (which includes uncommitted files) with a directory in the server; sshinto the server.- Start
tmuxand run the script. Among many things,tmuxlets you detach the session so you don’t have to wait for the job to finish before interacting with the server again.
When the script is sensible, I start diving into experiments with
multiple hyperparameter configurations.
A useful tool for this is
argparse.
It augments a Python script with commandline arguments, where you
add something like the following to your script:
parser = argparse.ArgumentParser()
parser.add_argument('--batch_size', type=int, default=128,
help='Minibatch during training')
parser.add_argument('--lr', type=float, default=1e-5,
help='Learning rate step-size')
args = parser.parse_args()
batch_size = args.batch_size
lr = args.lr
Then you can run terminal commands such as
python script1.py --batch_size=256 --lr=1e-4
This makes it easy to submit server jobs which vary these hyperparameters.
Finally, let’s talk about managing the output of experiments.
Recall the src/ directory structure above:
-- src/
-- checkpoints/
-- codebase/
-- log/
-- out/
-- script1.py
-- script2.py
We described the individual scripts and codebase/.
The other three directories are for organizing experiment output:
checkpoints/records saved model parameters during training. Usetf.train.Saverto save parameters as the algorithm runs every fixed number of iterations. This helps with running long experiments, where you might want to cut the experiment short and later restore the parameters. Each experiment outputs a subdirectory incheckpoints/with the convention,20170524_192314_batch_size_25_lr_1e-4/. The first number is the date (YYYYMMDD); the second is the timestamp (%H%M%S); and the rest is hyperparameters.log/records logs for visualizing learning. Each experiment belongs in a subdirectory with the same convention ascheckpoints/. One benefit of Edward is that for logging, you can simply pass an argument asinference.initialize(logdir='log/' + subdir). Default TensorFlow summaries are tracked which can then be visualized using TensorBoard (more on this next).out/records exploratory output after training finishes; for example, generated images or matplotlib plots. Each experiment belongs in a subdirectory with the same convention ascheckpoints/.
On data sets.
Data sets are used across many research projects. I prefer storing them
in the home directory ~/data.
On software containers. virtualenv is a must for managing Python dependencies and avoiding difficulties with system-wide Python installs. It’s particularly nice if you like to write Python 2/3-agnostic code. Docker containers are an even more powerful tool if you require more from your setup.
Exploration, Debugging, & Diagnostics
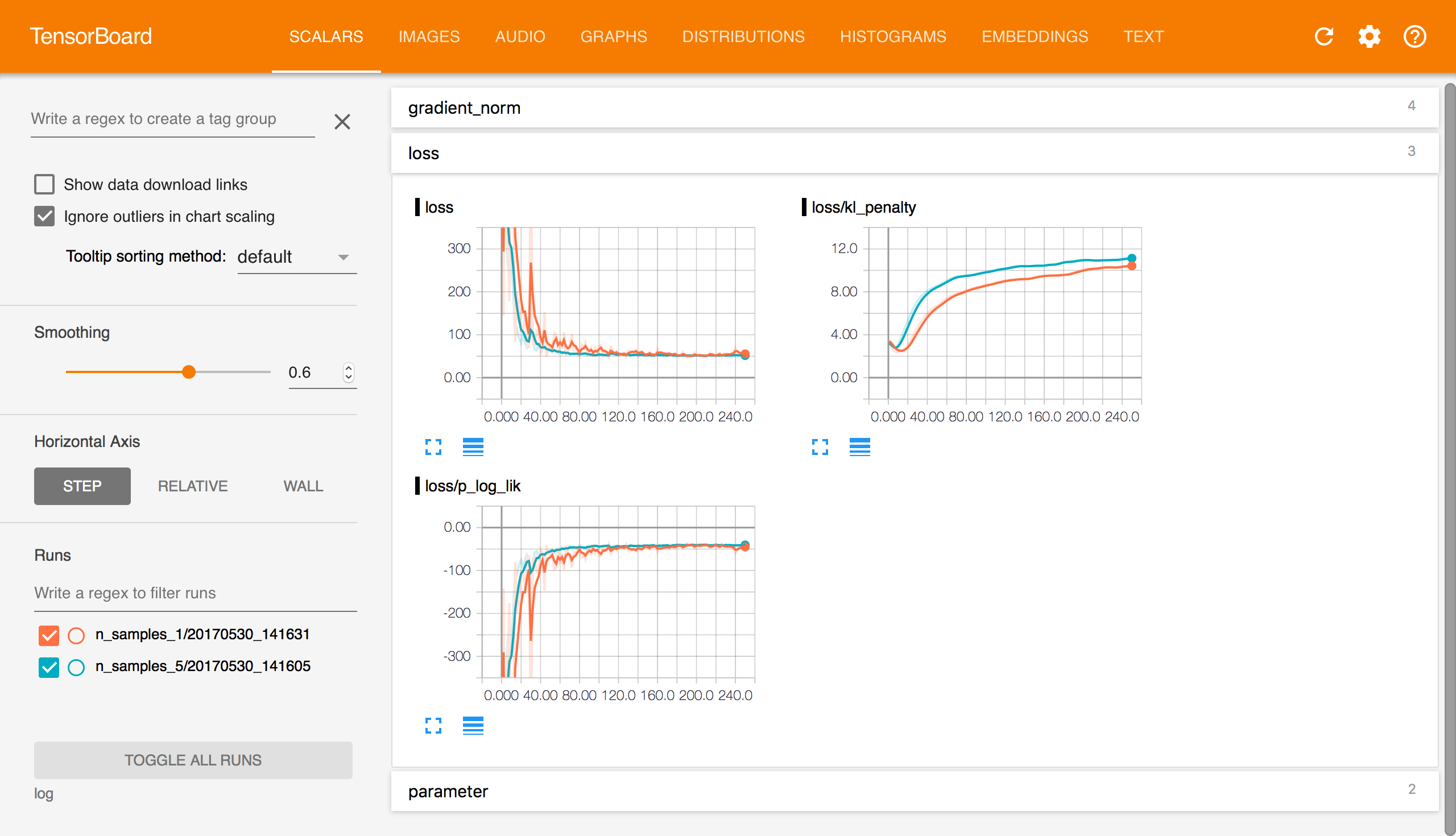
Tensorboard
is an excellent tool for visualizing and exploring your model
training. With TensorBoard’s interactivity, I find it
particularly convenient in that I don’t have to configure a bunch of
matplotlib functions to understand training. One only needs to percolate a
bunch of tf.summarys on tensors in the code.
Edward logs a bunch of summaries by default in order to visualize how
loss function values, gradients, and parameter change across
training iteration.
TensorBoard also includes wall time comparisons, and
a sufficiently decorated TensorFlow code base provides a nice
computational graph you can stare at.
For nuanced issues I can’t diagnose with TensorBoard specifically, I
just output things in the out/ directory and inspect those results.
Debugging error messages. My debugging workflow is terrible. I percolate print statements across my code and find errors by process of elimination. This is primitive. Although I haven’t tried it, I hear good things about TensorFlow’s debugger.
Improving Research Understanding
Interrogating your model, algorithm, and generally the learning process lets you better understand your work’s success and failure modes. This lets you go back to the drawing board, thinking deeply about the method and how it might be further improved. As the method indicates success, one can go from tackling simple toy configurations to increasingly large scale and high-dimensional problems.
From a higher level, this workflow is really about implementing the scientific method in the real world. No major ideas are necessarily discarded at each iteration of the experimental process, but rather, as in the ideal of science, you start with fundamentals and iteratively expand upon them as you have a stronger grasp of reality.
Experiments aren’t alone in this process either. Collaboration, communicating with experts from other fields, reading papers, working on both short and longer term ideas, and attending talks and conferences help broaden your perspective in finding the right problems and solving them.
Footnotes & References
1 This workflow is specifically for empirical research. Theory is a whole other can of worms, but some of these ideas still generalize.
2
The template for the master document is available
here.
3 There’s one caveat to Papers. I use it for everything: there are at least 2,000 papers stored in my account, and with quite a few dense textbooks. The application sifts through at least half a dozen gigabytes, and so it suffers from a few hiccups when reading/referencing back across many papers. I’m not sure if this is a bug or just inherent to me exploiting Papers almost too much.
- Neal, R. M. (1994). Bayesian Learning for Neural Networks (PhD thesis). University of Toronto.
- Pearl, J. (2000). Causality. Cambridge University Press.
- Wainwright, M. J., & Jordan, M. I. (2008). Graphical Models, Exponential Families, and Variational Inference. Foundations and Trends in Machine Learning, 1(1–2), 1–305.
